 +1 302-235-9792
+1 302-235-9792
OVERVIEW
Gene Universal offers CRISPR gene-edited cell line generation services based on a well-established gRNA/Cas9 ribonucleoprotein (RNP)–based CRISPR genome editing system. We provide customized cell line construction services for gene knockout, gene knock-in, and precise point mutation. The RNP-based CRISPR editing platform features high editing efficiency and reduced off-target effects.
SERVICE HIGHLIGHTS
-
 Expert Support
Expert Support
Our experienced CRISPR gene editing team provides customized cell line generation services tailored to specific research needs, including gene knockout, gene knock-in, and precise point mutations.
-
 Non-Viral Approach
Non-Viral Approach
Compared with traditional viral-based methods, our non-viral CRISPR editing strategy offers higher editing efficiency and reduced off-target effects.
-
 Rapid Turnaround
Rapid Turnaround
Gene Universal delivers a one-stop solution encompassing sgRNA synthesis, ssDNA synthesis, vector construction, and cell engineering, enabling fast and efficient project delivery.
SERVICE PROCEDURES
 Vector Construction
Vector Construction
· sgRNA or ssDNA synthesis
· Plasmid construction
 Gene Delivery
Gene Delivery
· Electroporation of RNP complexes
· Electroporation of RNP combined with ssDNA or plasmid
 Cell Line establishment
Cell Line establishment
· Antibiotic selection
· Monoclonal cell line establishment
 Genotype confirmation and QC
Genotype confirmation and QC
· PCR or qPCR
· Sanger sequencing
· WB Western blot (WB)
· Sterility and mycoplasma testing
SERVICE DETAILS
| Service Type | Service Name | Deliverables | Turnaround time(BD) | Price(USD) | Remarks |
|---|---|---|---|---|---|
| Knockout | Frameshift Mutation Knockout Cell Line (Sanger sequencing validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 60 | 2850 | RNP construction method |
| Knockout | Frameshift Mutation Knockout Cell Line (Sanger sequencing and Western blot Validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 70 | 3150 | RNP construction method |
| Knockout | Fragment Knockout Cell Line heterozygote(Sanger sequencing validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 60 | 4650 | RNP construction method |
| Knockout | Fragment Knockout Cell Line homozygote(Sanger sequencing validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 70 | 6900 | RNP construction method |
| Knockin | Point Mutation Knockin cell line heterozygote (Sanger sequencing validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 70 | 5700 | RNP construction method |
| Knockin | Point Mutation Knockin cell line homozygote (Sanger sequencing validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 70 | 8100 | RNP construction method |
| Knockin | Fragment Knockin cell line heterozygote (Sanger sequencing validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 80 | 6450 | RNP construction method |
| Knockin | Fragment Knockin cell line heterozygote(Sanger sequencing and Western blot Validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 90 | 6750 | RNP construction method |
| Knockin | Fragment Knockin cell line homozygote (Sanger sequencing validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 90 | 8700 | RNP construction method |
| Knockin | Fragment Knockin cell line homozygote(Sanger sequencing and Western blot Validation) | Target and Wild-type cells, >10^6 cells/tube *2 tubes for each type of cell | 100 | 9000 | RNP construction method |
Note: Cells, relevant detection antibodies, and any special culture media or special FBS must be provided by the customer.
CASE STUDY
-

The contributions indicate the inferred sequences present in the edited population and their relative proportions, in contrast to the indel distribution plot, which does not assign specific sequence contributions. Cut sites are shown as black vertical dotted lines, and the wild-type sequence is indicated by a “+” symbol on the far left.
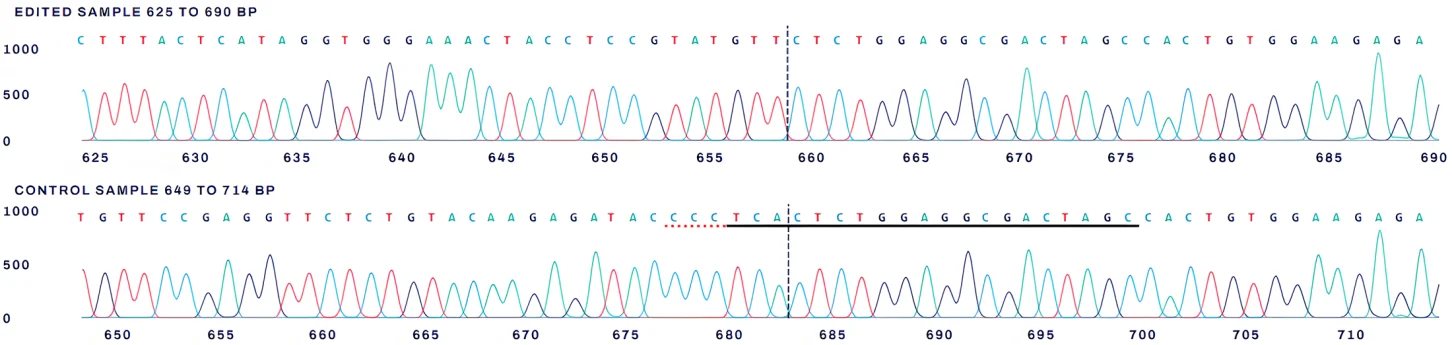
Sanger sequencing comparison of wild-type and edited samples around the guide RNA target site. The black underline indicates the guide sequence, the red underline marks the PAM site, and the dotted line denotes the expected cut site. Mixed base calls downstream of the cut site reflect error-prone repair following genome editing.
 44KD Target
44KD Target
 37KD GAPDH
37KD GAPDH
WT clone1 clone2 clone3
Figure: Validation of target gene knockout in T24 cells by Sanger sequencing and Western blot analysis.
Request A Quote

Download and fill out the order form below Order form

Please send e-mail to us for quotation sales@geneuniversal.com
Online Request Submission
Have a question about sales or support? Our team is ready to help!


